University of Waterloo researchers have developed a powerful new online tool that allows users to navigate through an interactive microbial tree of life, and to generate new scientific hypotheses and discoveries.
By integrating data across thousands of microbial genomes, “AnnoTree” provides a comprehensive framework for exploring the evolution of microbial genes and functions, and can be used to advance research across a wide range of industries including microbiology, biotechnology, industrial products, biofuels, and food science.
Increased understanding of gene function and diversity in microbes can lead to the identification of important biological phenomena, including the spread of antibiotic resistance, the evolution of bacterial pathogens, the origin of new gene families, and instances of gene transfer between specific bacterial genomes. AnnoTree is freely available at http://annotree.uwaterloo.ca.
A SCREENSHOT FROM 'ANNOTREE'
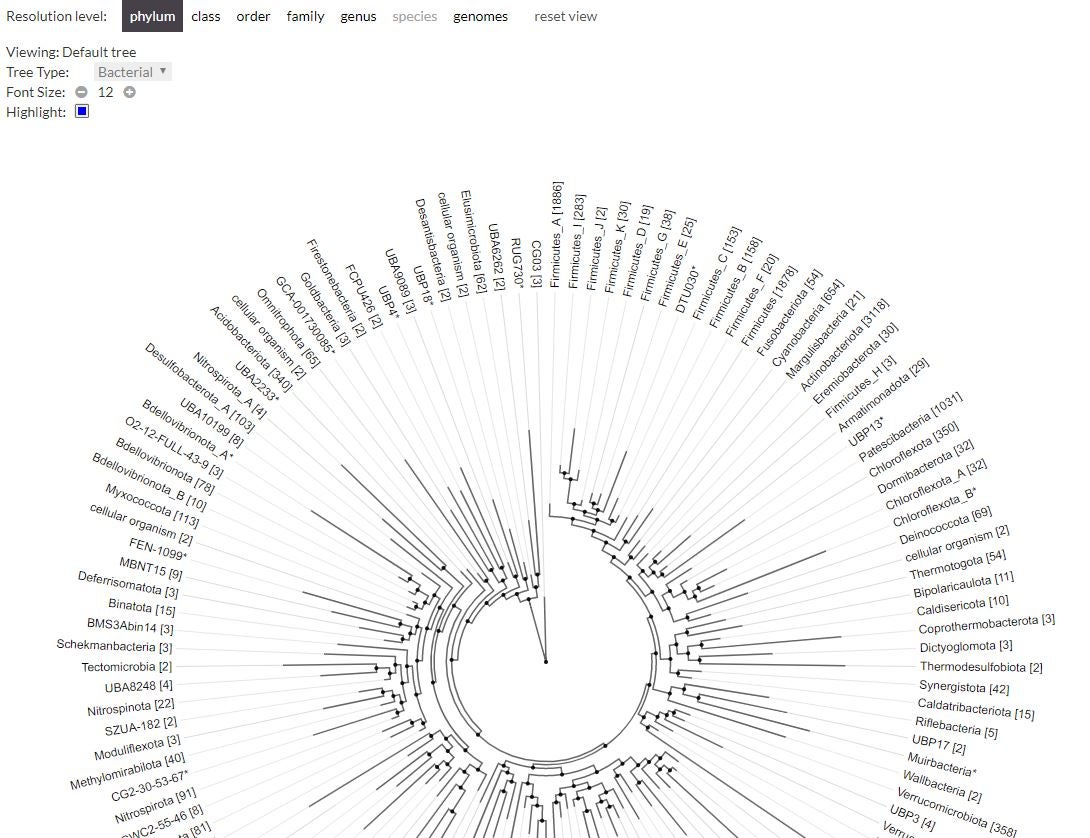
“Genomics
has
greatly
expanded
our
understanding
of
the
tree
of
life,
but
what
a
lot
of
researchers
ultimately
want
to
know
is
how
specific
genes
and
biological
functions
are
distributed
on
the
tree.
While
previously
this
type
of
analysis
could
take
a
substantial
amount
of
time
and
effort,
with
AnnoTree
it
takes
minutes
or
seconds,”
said
Prof.
Andrew
Doxey,
a
bioinformatician
at
the
University
of
Waterloo
and
member
of
the
Waterloo
Centre
for
Microbial
Research
(WCMR).
Through
AnnoTree’s
Web-based
interface,
users
may
query
any
gene
of
interest.
The
presence
or
absence
of
the
gene
will
then
be
“painted”
onto
the
microbial
tree
of
life
to
highlight
species
that
do
or
do
not
contain
the
gene.
Users
can
then
“zoom
in”
further
and
explore
specific
lineages
of
interest.
Users
may
download
publication-quality
images,
phylogenetic
trees,
gene
sequences,
and
other
information.
AnnoTree uses the recently developed Genome Taxonomy Database (GTDB), which provides a standardized, thorough description of the phylogeny and taxonomic nomenclature for over 27,000 Bacteria and 1,500 Archaea. All 28,941 microbial genomes in the GTDB were re-analyzed, and functional annotations were assigned to protein sequences during the development of AnnoTree. The resulting data is stored in an online database that is open and accessible to the research community. The AnnoTree database will be automatically updated to incorporate ongoing revisions to the GTDB taxonomy.
The AnnoTree application provides a reliable, automated tool for users to explore genes and functions of interest across microbes. The resulting genetic snapshots will enable researchers to quickly generate hypotheses, and identify genes and functions that are suitable for further study and application.
A paper on AnnoTree was recently published in the May 2019 issue of Nucleic Acids Research. Additional co-authors include Dr. Laura Hug (Biology, University of Waterloo), and Dr. Donovan Parks (Australian Centre for Ecogenomics).
Original post in Waterloo News.