 Radiomics, which involves the high-throughput extraction and analysis of a large amount of quantitative features from medical imaging data to characterize tumor phenotype in a quantitative manner, is ushering in a new era of imaging-driven quantitative personalized cancer decision support and management. While extremely powerful with the potential to make a tremendous clinical impact, the current state of radiomics rely entirely on existing, predefined imaging-based feature models based on aspects such as intensity, texture, and shape, which can greatly limit its ability to fully characterize the unique traits of different forms of cancer.
Radiomics, which involves the high-throughput extraction and analysis of a large amount of quantitative features from medical imaging data to characterize tumor phenotype in a quantitative manner, is ushering in a new era of imaging-driven quantitative personalized cancer decision support and management. While extremely powerful with the potential to make a tremendous clinical impact, the current state of radiomics rely entirely on existing, predefined imaging-based feature models based on aspects such as intensity, texture, and shape, which can greatly limit its ability to fully characterize the unique traits of different forms of cancer.
We have taken radiomics to the next level in personalized cancer quantification by introducing the concept of discovery radiomics, where we forgo the notion of predefined feature models by discovering customized, tailored radiomics feature models directly from the wealth of medical imaging data already available. This enables an unprecedented level of understanding and characterization of the unique cancer phenotype associated with different forms of cancer, allowing for the identification of a large amount of imaging-based features that capture highly unique tumor traits and characteristics beyond what can be captured using predefined feature models.
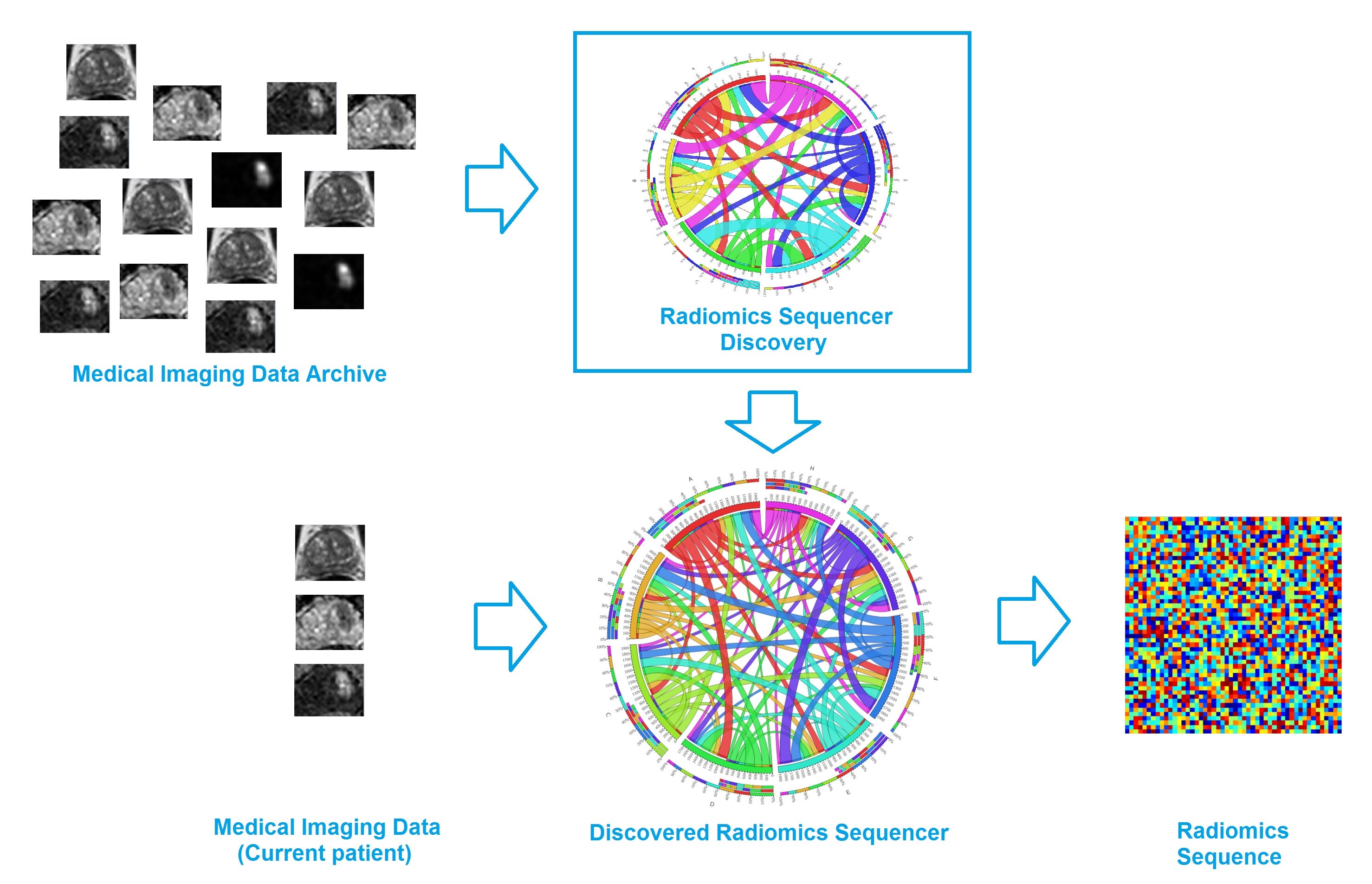
The discovery radiomics framework consists of the following steps (see Figure above). First, a wealth of standardized medical imaging data from past patients are fed into the radiomics sequencer discovery engine, where a customized radiomics sequencer is constructed based on a large number of radiomics features that were discovered to capture highly unique tumor traits and characteristics. Second, for a new patient case, the discovered radiomics sequencer is then used to extract a wealth of customized, tailored imaging-based features from the medical imaging data of the new patient case for comprehensive, custom quantification of the tumor phenotype.
Related people
Directors
Students
Alumni
Related publications
Shafiee, M. J., D. Kumar, F. Khalvati, M. A. Haider, and A. Wong,"Discovery Radiomics via StochasticNet Sequencers for Cancer Detection", CoRR, 2015.




