
What is a virus?
New research challenges conventional distinctions between viruses and cells
As more viruses are being discovered and studied, it’s becoming increasingly difficult to determine what distinguishes them from cells. New research conducted by Dr. Jozef Nissimov and his team has uncovered a genome of a cyanophage (a virus that infects cyanobacteria) that defies conventional distinctions between viruses and cells.
This work was conceived and led by Isaac Meza-Padilla, a graduate student in Nissimov’s lab who stumbled on the genome of this cyanophage, called PhiMa05, during his PhD research. His curiosity pushed him to follow up, leading to a discovery that could change models of virus-host evolution, infection strategies, and the role of phages in aquatic nutrient and protein synthesis dynamics.
Until recently, it was thought that ribosomes, the essential biological machines that synthesize proteins and bring to life the information contained in the genomes of organisms, were present only in cells and not in viruses. Even when viruses were later discovered with ribosomal proteins encoded in their genomes, there were never more than two or three.
The paper resulting from Meza-Padilla’s work, “A jumbo cyanophage encodes the most comprehensive ribosomal protein set in the known virosphere,” recently published in The ISME Journal, outlines how PhiMa05 challenges this assumption. With six ribosomal proteins and two proteins that participate in the formation of the ribosome, this is the first cyanophage discovered with ribosomal proteins and the virus with the most complete ribosomal protein set ever found.
After conducting evolutionary analyses, Meza-Padilla further discovered that an ancestor of PhiMa05 probably transferred its ribosomal protein genes to a non-photosynthetic cyanobacteria, and that the ribosomal genes acquired from the PhiMa05 ancestor were the only copies present in that cyanobacterial host. This indicates that these cellular organisms likely depend on the virus-derived ribosomal genes for protein synthesis and, consequently, life.
“The findings in this work support the idea that viruses actively reshape a host rather than just passively hijacking it for its own advantage,” says Nissimov.
This could change how we understand and deal with toxic algal blooms in lakes and rivers, since the productivity and turnover of dominant cyanobacteria in aquatic systems is affected by cyanophages. This is particularly relevant for PhiMa05, which infects cyanobacteria that form toxic freshwater algal blooms.
“This discovery implies a much deeper level of control over the host cell’s translational machinery than is typical for viruses,” says Meza-Padilla. “In aquatic biochemistry, this raises the possibility that some phages can fine-tune host physiology to boost viral replication under specific environmental conditions.”
And the implications of this work are possibly even more far-reaching. “This could be a major shift in our understanding of symbiosis, parasitism, and cellular dependence,” says Nissimov. “We tend to think of viruses as purely parasites and a nuisance, yet our findings suggest they can also be a kind of ‘evolutionary symbiont’ with the ability to move important genes from one species to another, in some cases helping cells to survive and replicate.”
Follow up research will further study this relationship between virus and cell, both experimentally in the lab and out in nature. “It would be fantastic to go on a field expedition to sample and study more PhiMa05-like viruses and their hosts," says Meza-Padilla. "There is a lot of exciting biology waiting to be discovered."
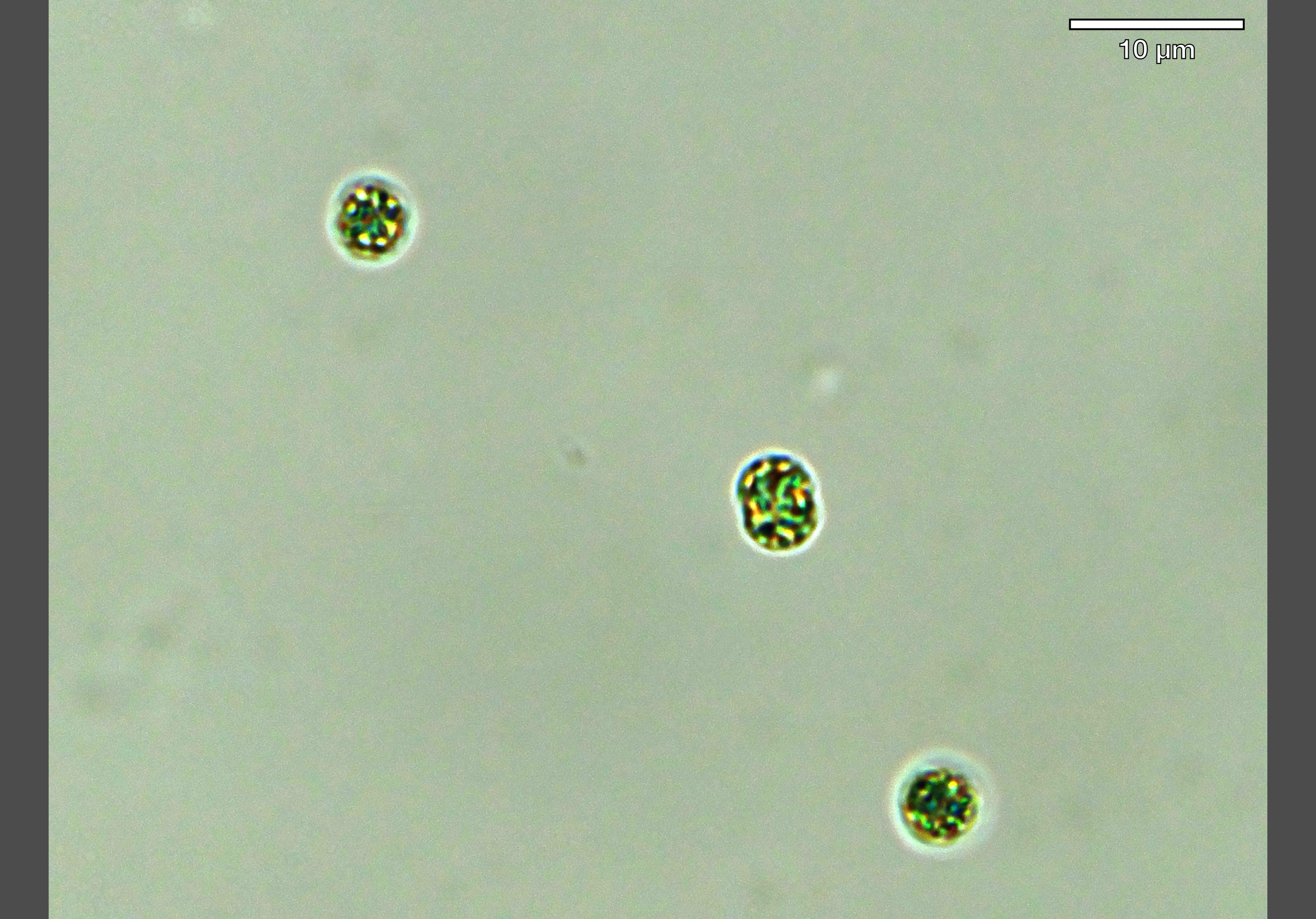
Microscopy image of toxic cyanobacterial cells in the lab, infected by a virus (courtesy of Isaac Meza-Padilla, PhD candidate).
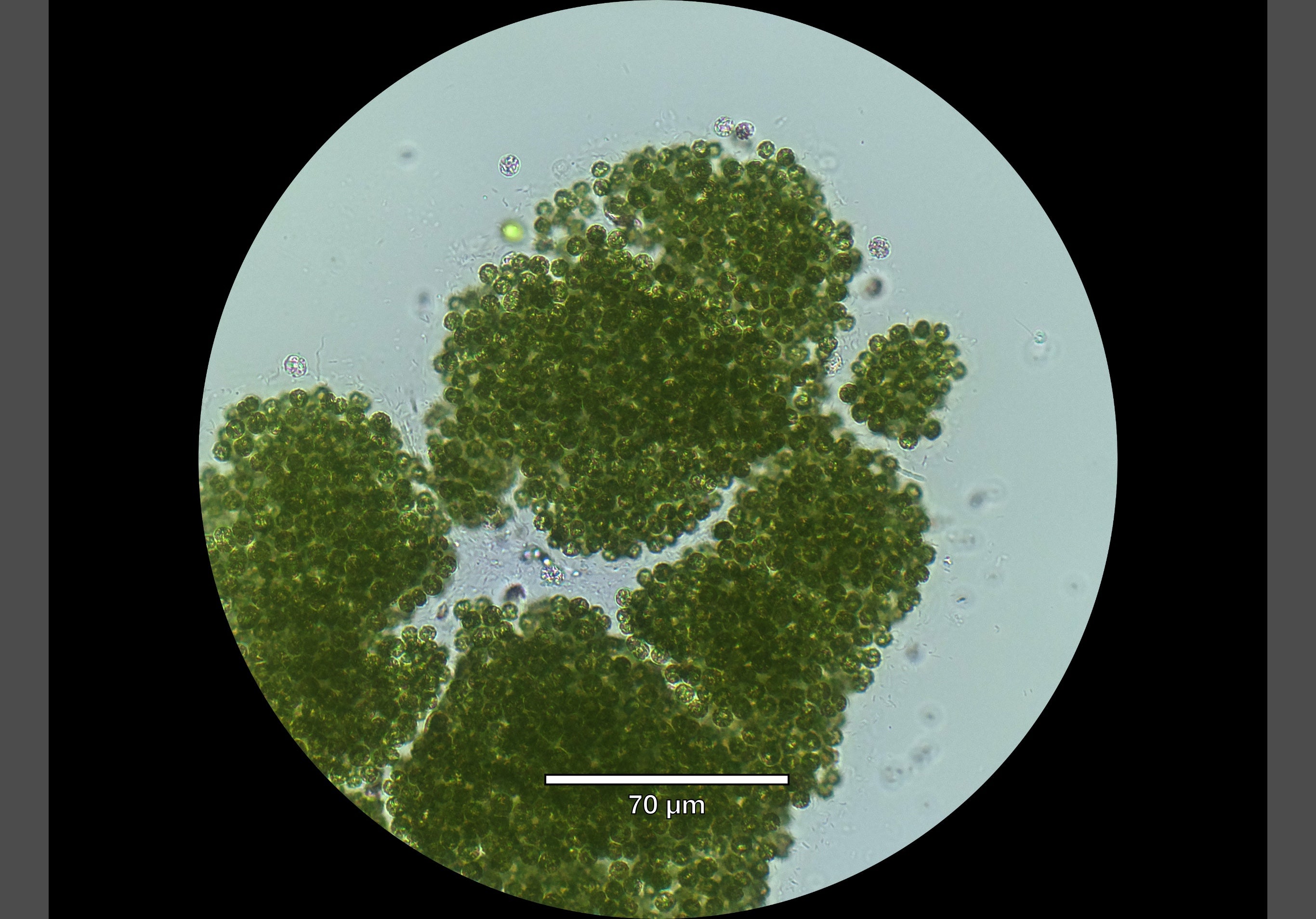

Microscopy image of toxic cyanobacterial colonies from Belwood Lake, September 2025 (courtesy of Cody Collis, PhD candidate).
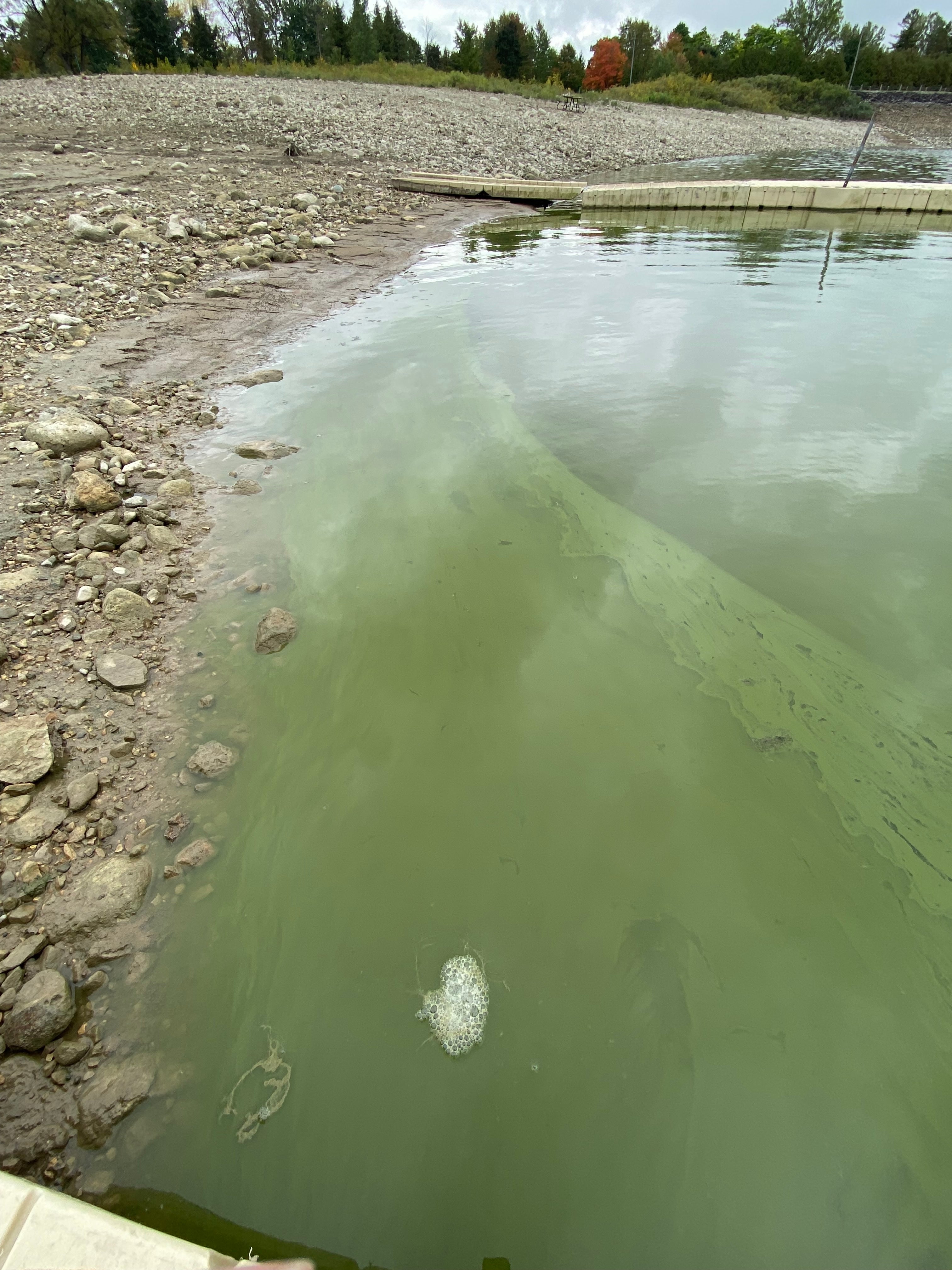

Toxic cyanobacterial bloom in Belwood Lake, September 2025 (courtesy of Cody Collis, PhD candidate).